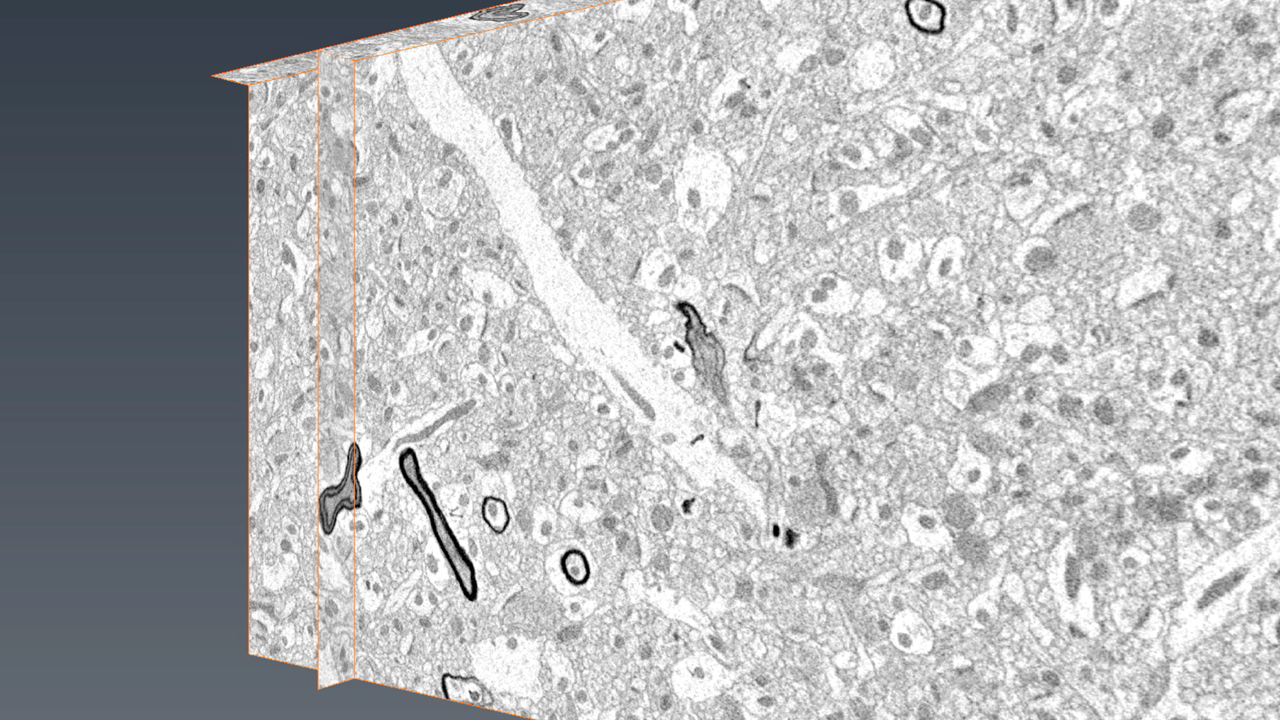
A 3D view of brain tissue imaged using a serial block face scanning electron microscope. Photo credit: Mayo Clinic Microscopy and Cell Analysis Core and Eugenia Trushina, Ph.D., Mayo Neurology
Before scientists can make breakthroughs in biomedical research, they must understand what’s going on in the body, often at a microscopic level.
But despite the range of sophisticated lab equipment often at scientists’ disposal, it can still be difficult to view some of our bodies’ tiniest structures at the right level of detail to see how their behaviors affect the body as a whole. The right level of microscopic imaging for observing how tissues, connections in the nervous system, cells, and cell parts all interact lands in a gap between what common lab microscopes can reach. Traditional optical microscopes can’t zoom in far enough, while more advanced electron microscopes see things at too tiny a level.
Recently, the University of Minnesota and Mayo Clinic teamed up to acquire an instrument that could bridge this divide and bring biomedical researchers at both institutions much-needed imaging capabilities. Through a collaboration between Mayo’s Microscopy and Cell Analysis Core and the U’s University Imaging Centers (UIC) and Research Computing, researchers will soon have access to a serial block face scanning electron microscope—a technology that can image the large, 3D data sets on cells and tissues that were, until now, unobtainable.
The new microscope is undergoing final installation and testing this week at its home on Mayo Clinic’s Rochester campus. Once installed it will drive forward biomedical research into a wide range of health problems, including neurodegenerative disease, cancer, diabetes, and developmental disorders.
Mark Sanders, UIC’s program director, said the microscope’s capabilities are hard to find elsewhere and can give researchers an edge in their studies.
“This system is the first of its kind in Minnesota and one of a handful of systems in the US,” Sanders said. “Together with Mayo Clinic, we are providing a unique level of imaging capability for researchers across the state.”
The collaboration doesn’t end with the imaging itself. Claudia Neuhauser, Ph.D., director of Research Computing, said researchers who use the microscope will also receive help drawing insights and meaning out of the resulting data.
“After using this new microscope to image a specimen, researchers will have an enormous amount of raw data that—until processed, analyzed, and interpreted—cannot play a meaningful role in their research,” Neuhauser said. “Our experts are here to assist in data analysis to improve reproducibility and expedite this stage of the research.”
The microscope’s purchase was supported by the state-funded Minnesota Partnership for Biotechnology and Medical Genomics, a collaboration between the U, Mayo Clinic, and the state of Minnesota designed to improve health, save lives, bolster Minnesota’s economy, and position the state as a world leader in biotechnology and biomedical research.
While the microscope will be housed on the Mayo campus in Rochester, it will also be available to U of M researchers and industry R&D with approved projects. Researchers can begin submitting requests to use the microscope for their projects now by contacting UIC.
Core Facilities Boost Imaging Capabilities
The serial block face scanning electron microscope joins a collection of cutting-edge technologies available to researchers through UIC, which is part of the U’s core research facilities.
Within the network of imaging facilities that make up UIC, two dozen advanced imaging systems are available to meet researchers’ needs in advanced optical imaging and basic electron microscopy. The centers help researchers design imaging experiments; choose a suitable imaging system; train on that system; and process, visualize, and analyze images.
Staff at UIC are also on hand to educate the University community about new imaging technologies, provide consulting and hands-on training for research projects, and assist with grant or publication writing.